plprot
- Plastids from etiolated rice tissue ["etioplast"]
|
Further details in:
Zychlinski et al., Molecular and Cellular Proteomics, 4:1072-1084, 2005.
|
Etioplasts are the endproduct of plastid differentiation in photosynthetic tissues
in the absence of light. They are characterized by a paracristalline prolamellar
body, that consists of a ternary complex of NADPH, protochlorophyllide oxidoreductase
(POR) and protochlorophyllide (Buchanan et al., 2000). Since etioplasts of higher
plants are unable to catalyze the conversion of protochlorophyllide to chloroplhyllide
in the absence of light, they are photosynthetically inactive. As a consequence,
their metabolism remains heterotrophic, i.e. they depend on the import of metabolites
from the cytosol to maintain their basic metabolic activities. Since etioplasts can
rapidly re-differentiate into photosynnthetically active chloroplasts upon illumination,
they were often used to study intial events in plastid differentiation processes.
Additionally, they are an ideal asset to analyze the plastid proteome in the absence
of highly-abundant photosynthetic proteins (Baginsky and Gruissem, 2004).
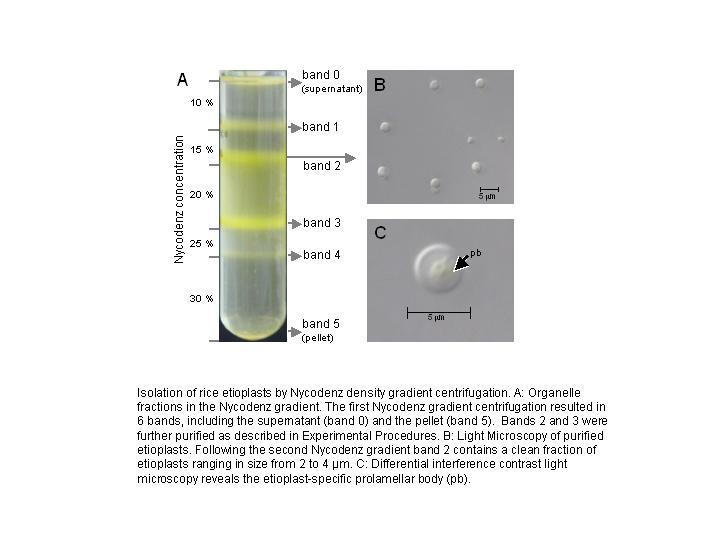
For our analysis, we have devised a new isolation strategy using a combination
of differential centrifugation and Nycodenz density gradient centrifugation
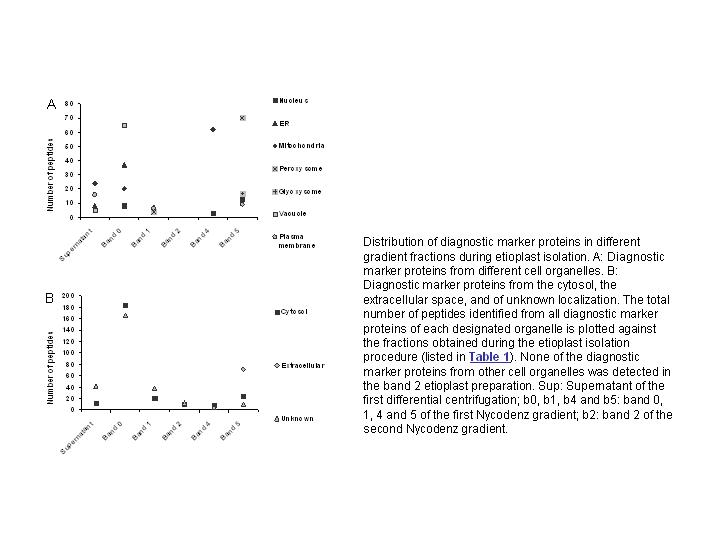
(details in Zychlinski et al., 2005). A thorough analysis of cellular marker
proteins from other cell organelles in the Nycodenz gradient suggested, that
the Nycodenz density gradient centrifugation resulted in highly pure etioplasts
that were efficiently separated from all other cell organelles (FIGURE 1, FIGURE 2).
All identified marker proteins from the Nycodenz gradient fractions are provided
in Table 1. Using highly purified etioplasts for the proteome analysis, we
identified 240 different proteins by LC-ESI-MS/MS analysis (Zychlinski et al., 2005).
A surprisingly large number of these proteins is involved in gene expression
suggesting that etioplasts are a special plastid type that is prone to initiate
adaptations of gene expression rates immediately upon receipt of a light signal.
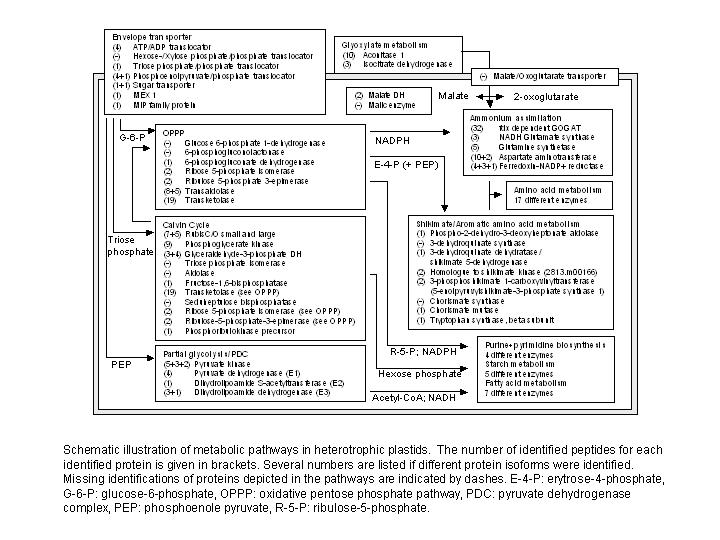
Identified proteins from characteristic heterotrophic metabolic functions are
assembled in FIGURE 3. All identified proteins can be searched in plprot, either
by key word or by BLAST search.
|

Top
|
Top
|
Top
|
|